library(here)
library(tidyverse)Exploration
flu <- readRDS(here("fluanalysis/data/processed_data/flu.rds"))Let’s take a look at some summary statistics for BodyTemp and Nausea
summary(flu$BodyTemp) Min. 1st Qu. Median Mean 3rd Qu. Max.
97.20 98.20 98.50 98.94 99.30 103.10 summary(flu$Nausea) No Yes
475 255 Next, let’s take a look at this distribution of BodyTemp.
flu %>% ggplot(aes(x = BodyTemp)) + geom_histogram(bins = 20)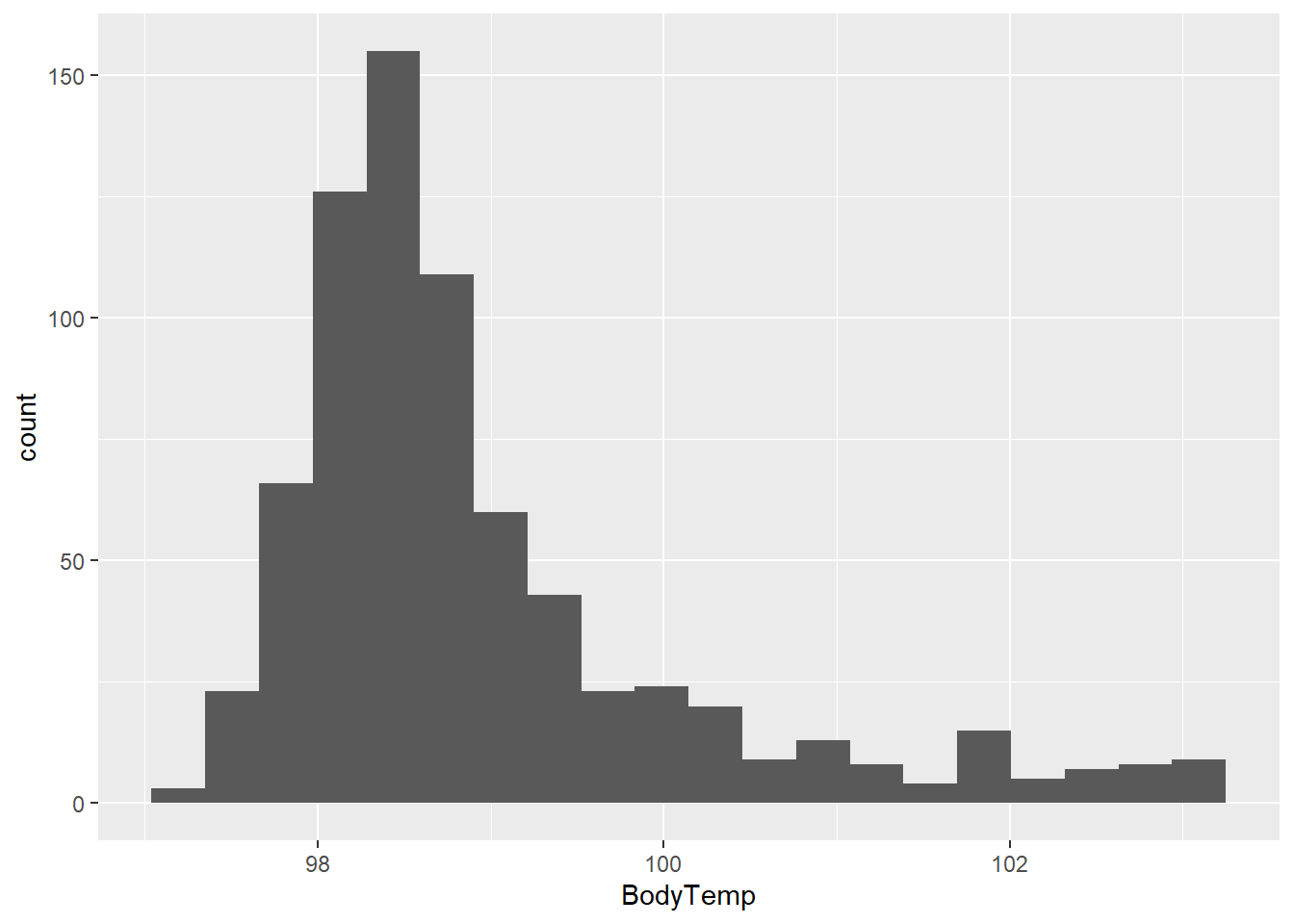
I’m also interested in the intensity variables… Let’s make a box plot of a couple of these with BodyTemp.
flu %>% ggplot(aes(x= BodyTemp, y = Myalgia, color = Myalgia)) +
geom_boxplot() 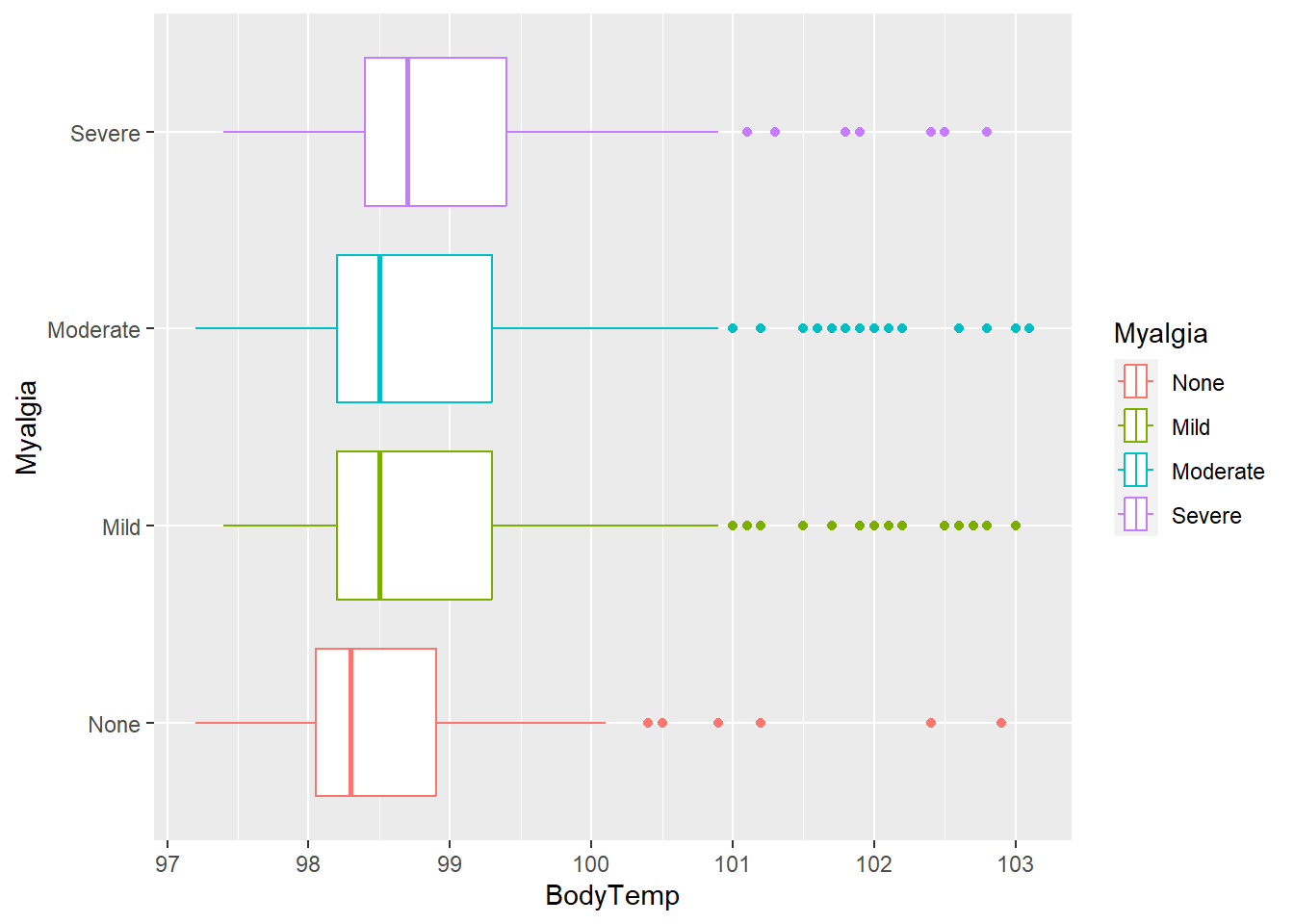
flu %>% ggplot(aes(x= BodyTemp, y = Weakness, color = Weakness)) +
geom_boxplot()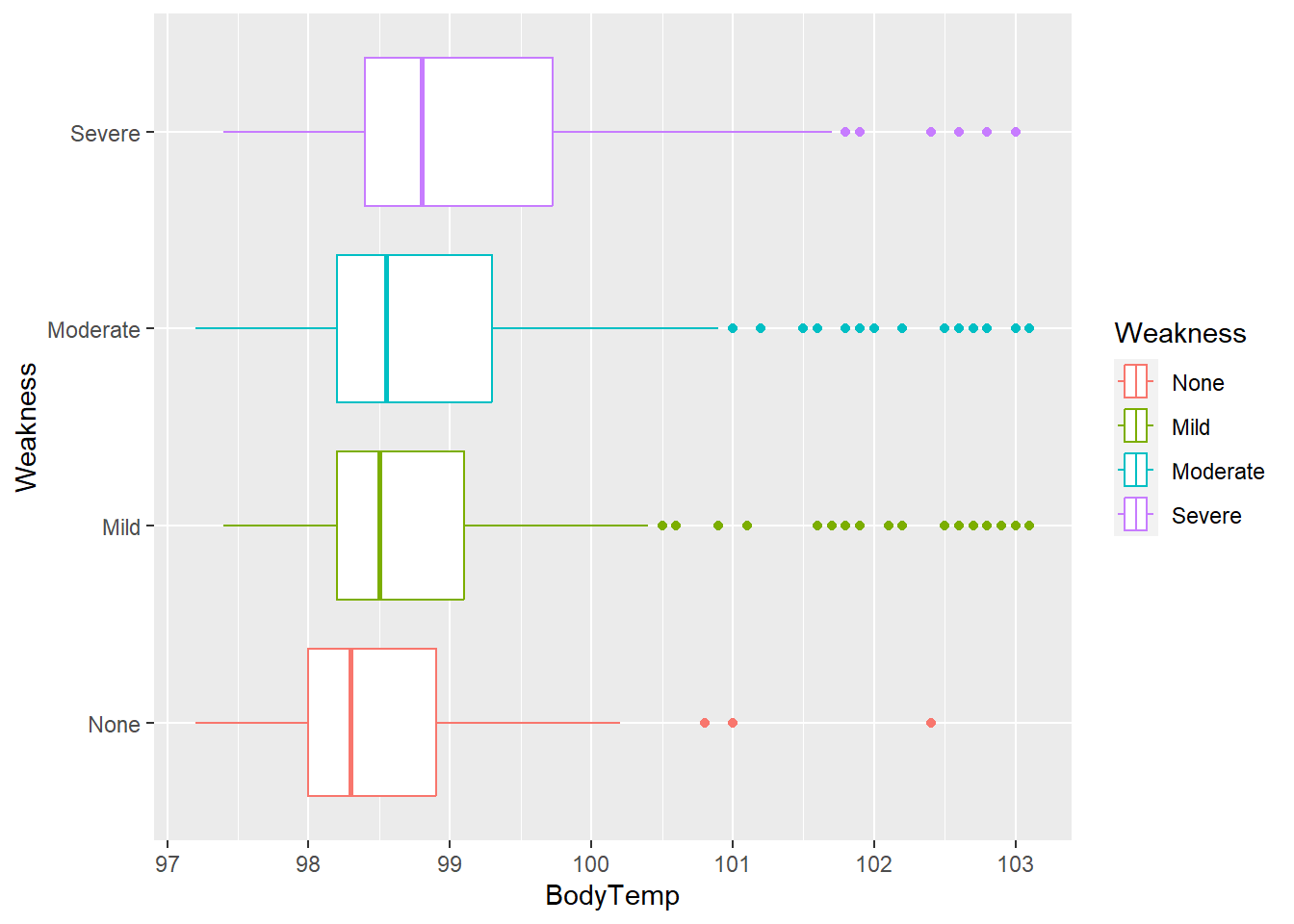
Median body temperature appears to increase with increasing intensity of myalgia/ weakness.
Let’s look at this as a histogram for weakness.
flu %>% ggplot(aes(x = BodyTemp, fill = Weakness)) +
geom_histogram(bins = 20) 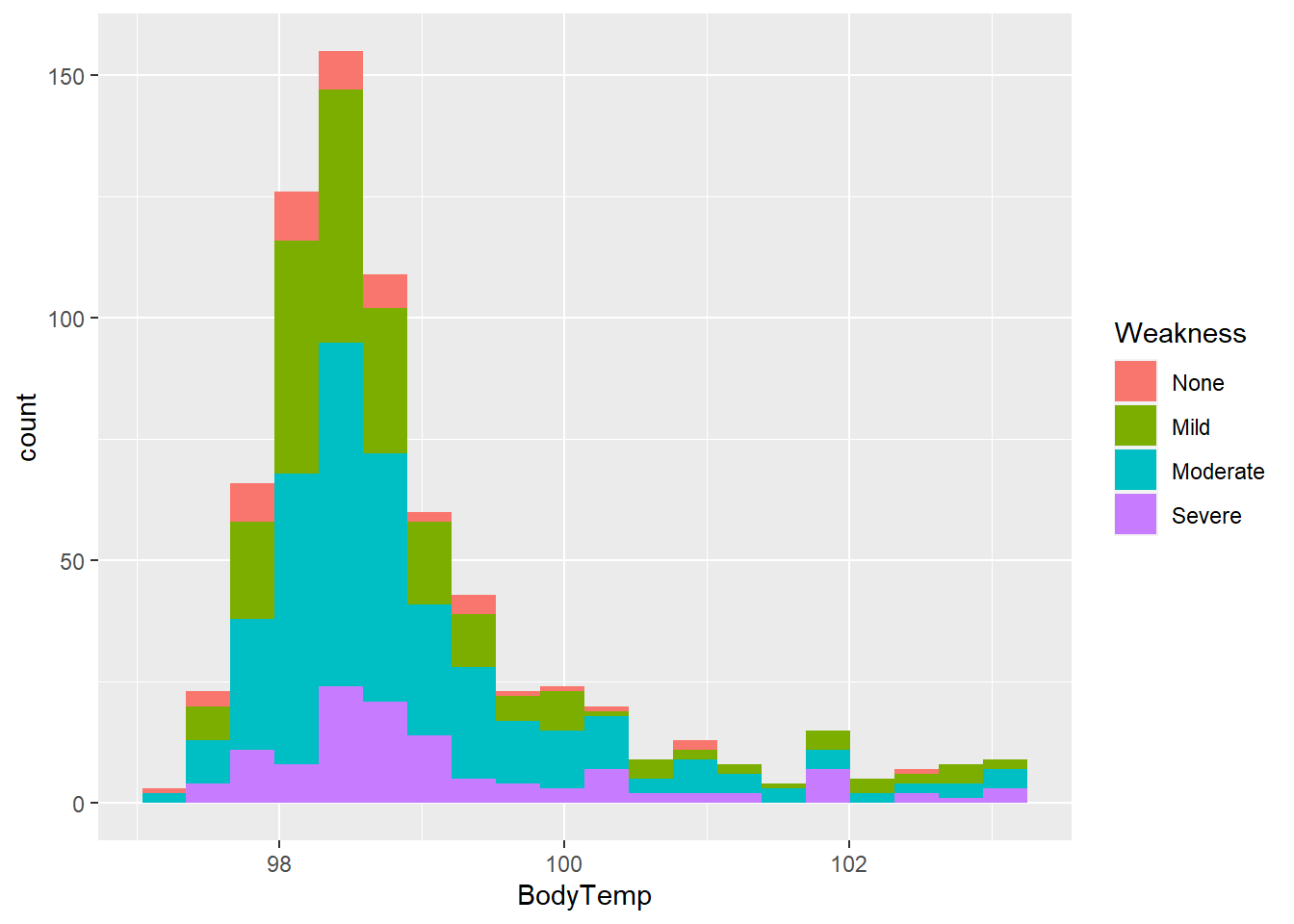
Weakness by Nausea contingency table
table(flu$Weakness,flu$Nausea)
No Yes
None 39 10
Mild 172 51
Moderate 210 128
Severe 54 66Myalgia by Nausea contingency table
table(flu$Myalgia,flu$Nausea)
No Yes
None 63 16
Mild 159 54
Moderate 198 127
Severe 55 58Cough Intensity by Nausea contingency table
table(flu$CoughIntensity,flu$Nausea)
No Yes
None 30 17
Mild 99 55
Moderate 232 125
Severe 114 58Now let’s take visualize this.
flu %>% ggplot(aes(x= Weakness, fill = CoughIntensity)) +
geom_histogram(stat="count")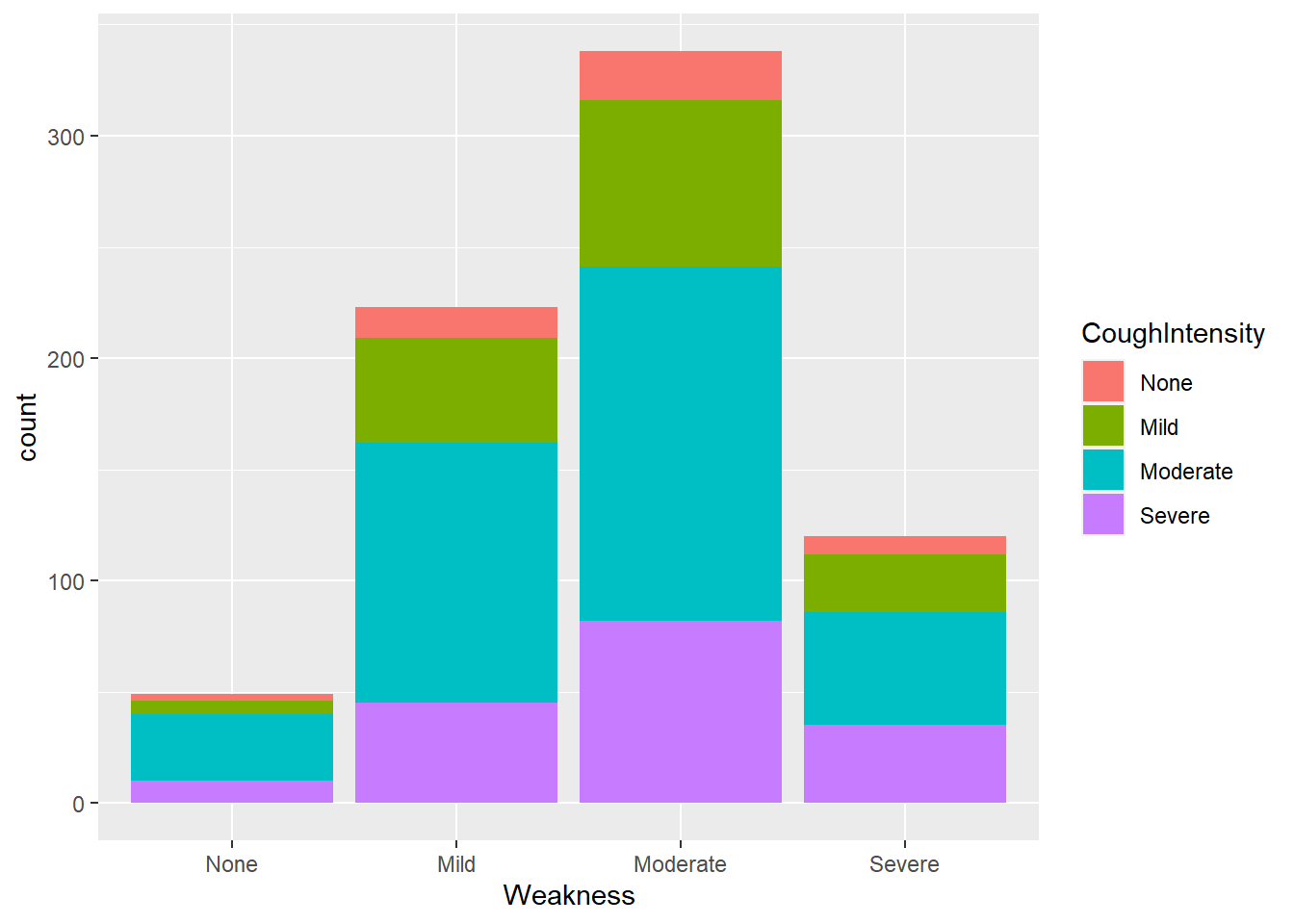
flu %>% ggplot(aes(x= Weakness, fill = Myalgia)) +
geom_histogram(stat="count")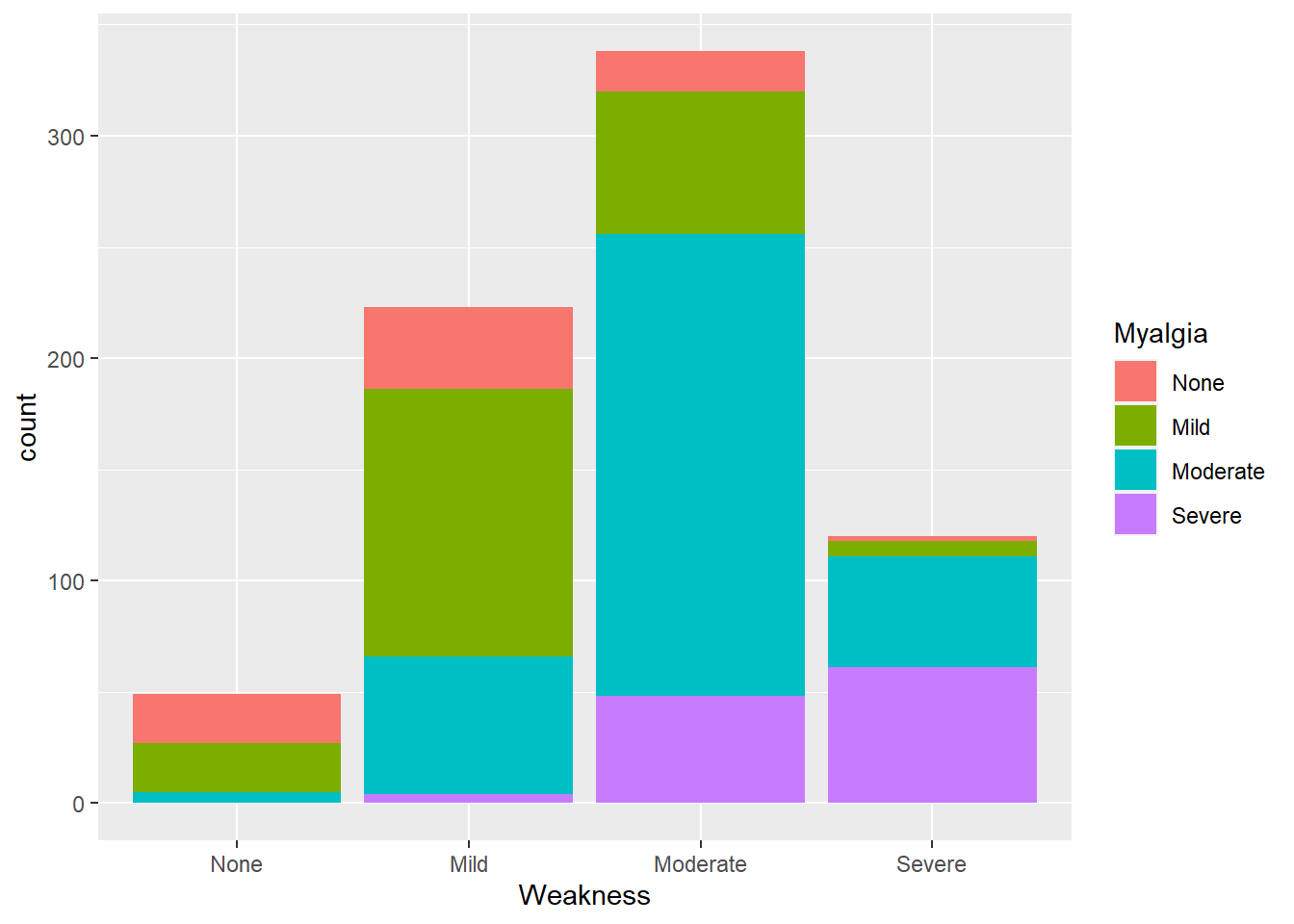
flu %>% ggplot(aes(x= Weakness, fill = Nausea)) +
geom_histogram(stat="count")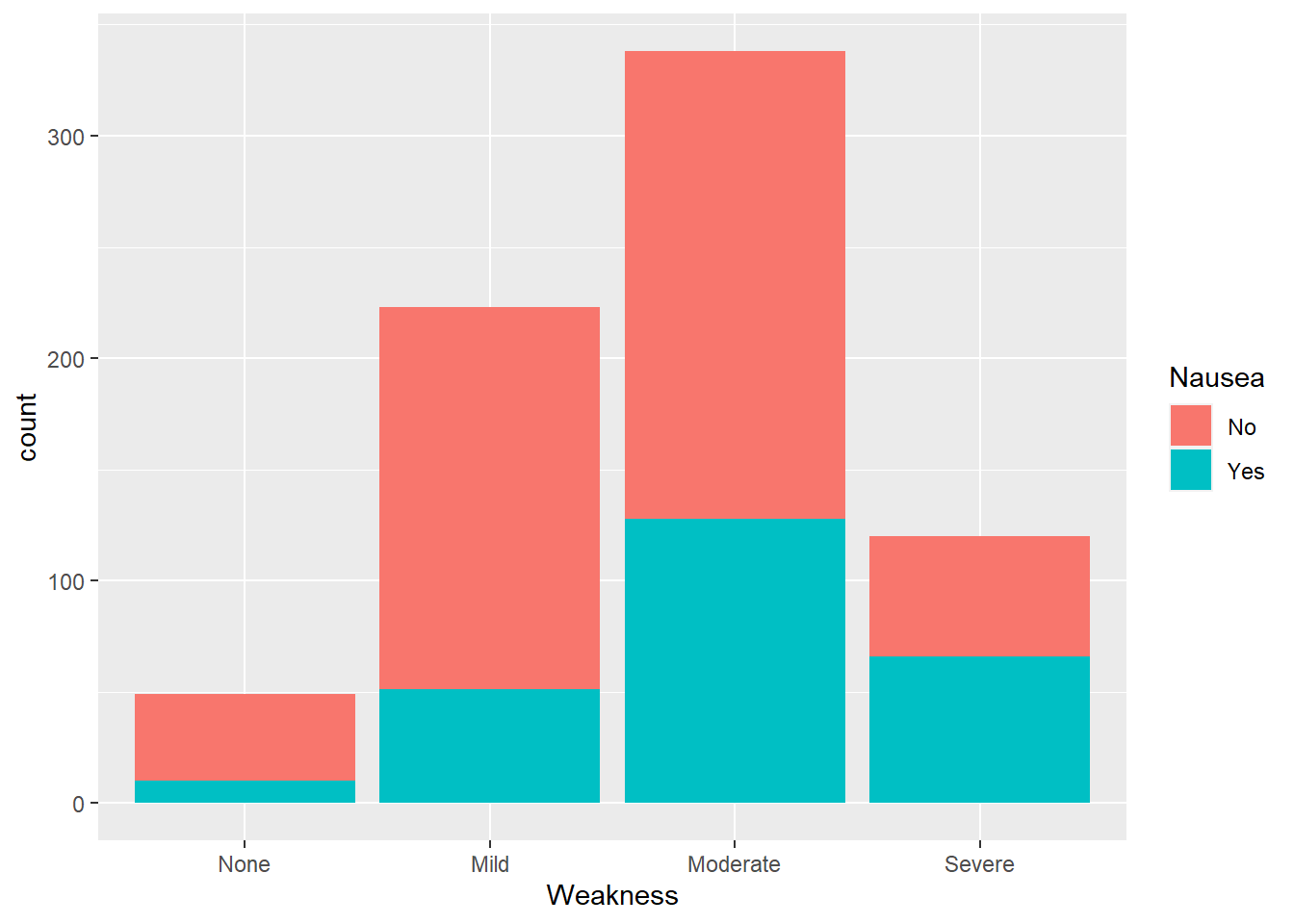
Weakness by Nausea contingency table
table(flu$Weakness,flu$Myalgia)
None Mild Moderate Severe
None 22 22 5 0
Mild 37 120 62 4
Moderate 18 64 208 48
Severe 2 7 50 61